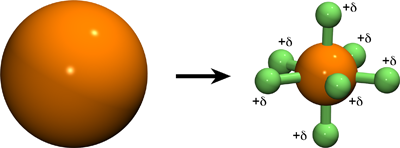
This page describes how to implement out multisite cation models as described in:
A. Saxena and D. Sept, Multisite ion models that improve coordination and free energy calculations in molecular dynamics simulations, J. Chem. Theor. Comp. 9(8) 3538-42, 2013.
Note that these instructions describe the implementation in Amber, but other implementations have been performed successfully and are discussed below. Also note that these procedures and files differ slightly from the supplemental materials published with the manuscript, however, these should be simpler and faster.
- First, replace the spherical ion with the multisite ion structure from either the calcium or magnesium pdb files.
- Use xleap to generate *parm (topology) and *crd (structure) files for the system. Addition of water and/or counterion can also be done at this step. Also, make sure to load the *lib files (cal.lib and mag.lib) and *frcmod files (frcmod.mca and frcmod_mg.mmg). This must be done so that the new atom types are recognized. N.B. that the frcmod and cal.lib files are one of the changes mentioned above.
- As input, Amber uses the r and ε notation, but since our dummy atoms have only a repulsive part of the potential and no attractive part, there is no way to make this work mathematically. However, when xleap generates the topology file, it converts this back to the A/B form that we discuss in the paper. The parameters are correct for the central atom (XZ) and the A parameters are correct for the dummy atoms, however, the B values are non-zero and need to be fixed. To do this, we provide a simple python script (zerobvalues.py). This script will go through the topology file generated above, locate the dummy atoms, determine the dummy atom type, and correct the LENNARD_JONES_B parameters.
Example of Calcium in Water >tleap -s -f leaprc.ff99SB > loadoff bondmiss_cal.lib Loading library: ./bondmiss_cal.lib > loadamberparams frcmod_ca.mca Loading parameters: ./frcmod_ca.mca Reading force field modification type file (frcmod) Reading title: \# parameters for multisite Ca++ from Saxena and Sept JCTC (2013) > mol = loadpdb calcium_ms.pdb Loading PDB file: ./calcium_ms.pdb total atoms in file: 8 > solvatebox mol TIP3PBOX 10.0 Solute vdw bounding box: 2.712 2.628 2.800 Total bounding box for atom centers: 22.712 22.628 22.800 Solvent unit box: 18.774 18.774 18.774 Total vdw box size: 25.763 25.372 25.855 angstroms. Volume: 16900.198 A^3 Total mass 6057.424 amu, Density 0.595 g/cc Added 334 residues. > addions2 mol Cl- 2 Adding 2 counter ions to "mol" using 1A grid Grid extends from solute vdw + 2.47 to 8.24 Resolution: 1.00 Angstrom. grid build: 0 sec Calculating grid charges charges: 0 sec Placed Cl- in mol at (12.65, 7.32, -1.54). Placed Cl- in mol at (-13.35, -2.68, 4.46). Done adding ions. > saveamberparm mol Ca_Model_bond.top Ca_Model_bond.crd >quit >./zeroBvalues.py This is a simple script designed to zero out the Lennard-Jones B values for dummy atoms in our multisite ion model. Please input topology file: Ca_Model_bond.top Modified parameter file will be Ca_Model_bond.top.mod Topology file has 1012 atoms and 5 atom types D1 atom at position 7 in atom list D1 atom has type 2 Making substitution in LENNARD_JONES_BCOEF at positions: [2, 3, 5, 8, 12]
Implementation in Gromacs
This is under development.
